 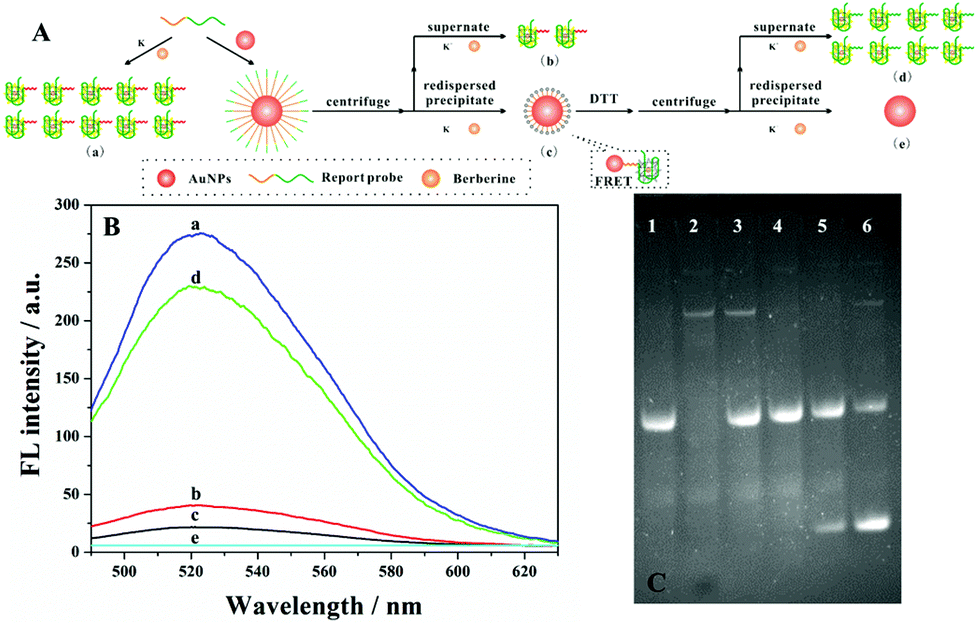
Sequencing of bacterial genomes also prompted the development of bioinformatics and software required to handle a large amount of data.Īlthough dideoxy chain terminator sequencing was absolutely critical in the development of modern microbiology, from molecular to environmental sciences, the method itself has been relatively troublesome. Those early years were crucial not only for gaining basic knowledge about bacterial genomes, but also to develop sequencing strategies and lay the foundation for future microbial genomics and transcriptomics. 1995) A decade between the mid-1990s and mid-2000s gave us genomic sequences of other bacterial species, and we saw the rise of comparative genomics and metagenomics (for a review, see Loman and Pallen ( 2015)). 1981)-and years later first complete bacterial genome of Haemophilus influenzae (Fleischmann et al. The technology was used to sequence in 1981 first part of the human genome-mitochondrial DNA (Anderson et al. 1986).įor over 30 years, “Sanger sequencing” allowed to unveil crucial secrets of life. Only later development of fluorescence-based methods and automated instrumentation allowed to increase speed and the scale of dideoxy chain terminator sequencing (Smith et al. In addition, gels needed exposure to photographic film and manual sequence reading. This first genome sequencing of bacteriophage φX174 was performed using long sequencing gels and radioactively labeled nucleotides. The major challenge in understanding genomic and transcriptomic data lies today in combining it with other sources of global data such as proteome and metabolome, which hopefully will lead to the reconstruction of regulatory networks within bacterial cells that allow communicating with the environment (signalome and interactome) and virtual cell reconstruction.Ī little over 40 years ago, Fred Sanger published his paper describing dideoxy chain terminator sequencing (Sanger et al. Finally, NGS is a key technique to study the organization of the bacterial life-from complex communities to single cells. The use of RNA sequencing techniques allows studying in detail the complex regulatory processes in the bacterial cells. Today, the use of NGS expended capabilities of diagnostic microbiology and epidemiology. NGS started its infancy from basic structural and functional genomics, to mature into the molecular taxonomy, phylogenetic and advanced comparative genomics. The major goal of next-generation sequencing for a microbiologist is not really resolving another circular genomic sequence. This review will discuss applications of high-throughput methods to study bacteria in a much broader context than simply their genomes. Human Genome Project was a huge impulse to improve sequencing technologies, and unprecedented financial and human effort prompted the development of cheaper high-throughput technologies and strategies called next-generation sequencing (NGS) or whole genome sequencing (WGS). However, this method of sequencing has been time-consuming, laborious and remains expensive even today. For the last 40 years, “Sanger sequencing” allowed to unveil crucial secrets of life.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
